Tetrahedra microFE pipeline - non-linear simulation in Calculix
Created on: 23.05.2021
Last update: 14.01.2023
By: Gianluca Iori, 2023
The specimen scanned for this example is courtesy of Prof. Dr.-Ing. Claudia Fleck, Institut für Werkstoffwissenschaften und technologien, TU Berlin
For more info on the dataset see the paper: Kaya et al. (2019), Foams of Gray Cast Iron as Efficient Energy Absorption Structures: A Feasibility Study.
Adv. Eng. Mater., 21: 1900080. https://doi.org/10.1002/adem.201900080Code license: MIT
Narrative license: CC-BY-NC-SA
Command line execution
The pipeline can be executed from the command line using the ciclope command:
python ciclope.py input.tif output.inp -vs 0.0065 0.0065 0.0065 --smooth -r 1.2 -t 90 --vol_mesh --tetrafe --template input_templates/tmp_example02_tens_Nlgeom_steel.inp -v
For info on the solver see the Calculix homepage

Configuration and imports
import sys
sys.path.append('./../../')
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import mcubes
from scipy import ndimage, misc
from skimage.filters import threshold_otsu, gaussian
from skimage import measure, morphology
from ciclope.utils.recon_utils import read_tiff_stack, plot_midplanes
from ciclope.utils.preprocess import remove_unconnected
import ciclope
matplotlib.rcParams['figure.dpi'] = 300
font = {'weight' : 'normal',
'size' : 8}
plt.rc('font', **font)
astropy module not found
Uncomment and run the following cell for visualizations using itkwidgets
# import vtk
# import itk
# from itkwidgets import view
ccx2paraview, dat2txt, and pandas are needed for the postprocessing of results
import ccx2paraview
import dat2txt
import pandas as pd
%%html
<style>
table {float:left}
</style>
Load input data
input_file = './../../test_data/steel_foam/rec_8bit_0_rescale_025/B_matrix_01_0000.tif'
Input data description
Scan parameters |
|
|---|---|
Beamline |
TOMCAT@SLS |
Sample |
Sintered steel foam strut from Kaya_2016 |
Energy |
34 keV |
Camera |
PCO.edge5 |
Optics |
4x |
Voxel size |
1.625 micron |
Preliminary operations |
downsampled (5X; quadratic interpolation) |
data_3D = read_tiff_stack(input_file)
vs = np.ones(3)*1.625e-3*4 # [mm]
data_3D = data_3D[100:300, 100:300, 100:300]
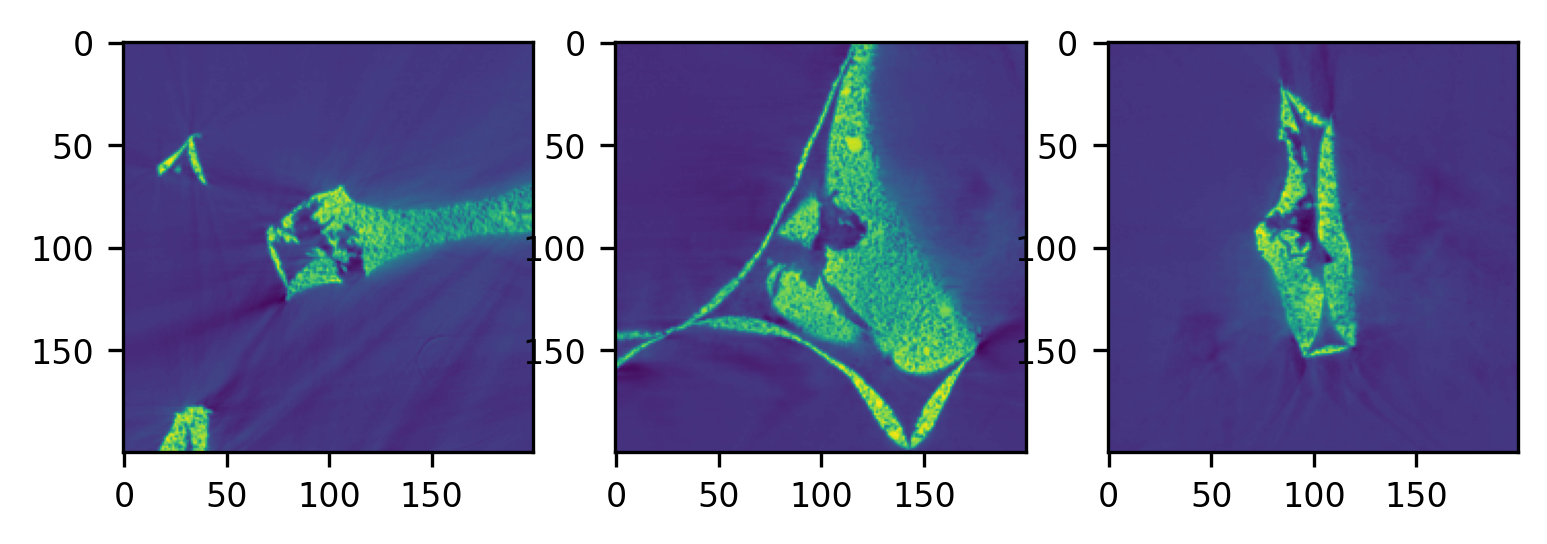
# Inspect dataset
plot_midplanes(data_3D)
plt.show()
Inspect the dataset with itkwidgets
# viewer = view(data_3D, ui_collapsed=True)
# viewer.interpolation = False
Launch the itk viewer
# viewer
Pre-processing
Gaussian smooth (optional)
data_3D = gaussian(data_3D, sigma=1, preserve_range=True)
Resize (optional)
resampling = 1.2
# resize the 3D data using spline interpolation of order 2
data_3D = ndimage.zoom(data_3D, 1/resampling, output=None, order=2)
# correct voxelsize
vs = vs * resampling
# Inspect again the dataset
plot_midplanes(data_3D)
plt.show()

Thresholding
# use Otsu's method
T = threshold_otsu(data_3D)
print("Threshold: {}".format(T))
Threshold: 88
# plot the input image histogram
fig2, ax2 = plt.subplots()
plt.hist(data_3D.ravel(), bins=40)
plt.show()

# apply the threshold
# BW = data_3D > T
BW = data_3D > 90
Morphological close of binary image
BW = morphology.closing(BW, morphology.cube(5))
Detect largest isolated cluster of voxels
Label the BW 3D image
# [labels, n_labels] = measure.label(BW, None, True)
[labels, n_labels] = measure.label(BW, None, True, 1) # 1 connectivity hop
Inspect the labels with napari
# import napari
# viewer = napari.view_image(labels)
# count occurrences of each label label
occurrences = np.bincount(labels.reshape(labels.size))
# find largest unconnected label
largest_label_id = occurrences[1:].argmax()+1
L = labels == largest_label_id
You can do the same using the function remove_unconnected contained in preprocess
L = remove_unconnected(BW)
# Inspect dataset
# plot_midplanes(np.bitwise_xor(L, BW))
plot_midplanes(L)
plt.show()

Generate meshes for visualization
Create triangles mesh of the outer model shell (optional.. expensive!)
# vertices, triangles = ciclope.shell_mesh(L, method='pymcubes')
Write VTK mesh with meshio
# meshio.write_points_cells("pippo_shell.vtk", vertices.tolist(), [("triangle", triangles.tolist())])
Create tetrahedra mesh
Volume meshing using CGAL through pygalmesh
filename_mesh_out = '/home/gianthk/PycharmProjects/CT2FE/test_data/steel_foam/B_matrix_mesh.vtk'
# import pygalmesh
# mesh = pygalmesh.generate_from_array(np.transpose(L, [2, 1, 0]).astype('uint8'), tuple(vs), max_facet_distance=1.2*min(vs), max_cell_circumradius=5*1.2*min(vs))
Alternatively use the method ciclope.tetraFE.cgal_mesh
For some reason this works only from script
mesh = ciclope.tetraFE.cgal_mesh(L, vs, 'tetra', 1.2*min(vs), 5*1.2*min(vs))
mesh.write(filename_mesh_out)
Visualize the mesh with itkwidgtes
# from itkwidgets import view
# reader = vtk.vtkUnstructuredGridReader()
# reader.SetFileName(filename_mesh_out)
# reader.Update()
# grid = reader.GetOutput()
# view(geometries=grid)
Write Abaqus input FE files
Generate tetrahedra-FE model with constant material properties
The method ciclope.tetraFE.mesh2tetrafe assumes that the material property definition is contained in the FE analysis .INP template file.
input_template = "./../../input_templates/tmp_example02_tens_static_steel.inp"
Inspect input template file
!cat {input_template}
** ----------------------------------------------------
** Material definition:
*MATERIAL, NAME=STEEL_4301
*ELASTIC
210000, 0.3333333333, 0
*DENSITY
7.85e-9
*SOLID SECTION, ELSET=SET1, MATERIAL=STEEL_4301
** ---------------------------------------------------
** Analysis definition. The Step module defines the analysis steps and associated output requests.
** More info at:
** https://abaqus-docs.mit.edu/2017/English/SIMACAECAERefMap/simacae-m-Sim-sb.htm#simacae-m-Sim-sb
** ---------------------------------------------------
*STEP
*STATIC
** All displacements fixed at bottom:
*BOUNDARY
NODES_B, 1, 3, 0.0
** Vertical displacement imposed at top:
*BOUNDARY
NODES_T, 3, 3, +0.1
** ---------------------------------------------------
** Output request:
*NODE FILE, OUTPUT=2D
U
*NODE PRINT, TOTALS=ONLY, NSET=NODES_B
RF
*EL FILE, OUTPUT=2D
S
*END STEP
filename_out = './../../test_data/steel_foam/B_matrix_tetraFE.inp'
Generate CalculiX FE input file
ciclope.tetraFE.mesh2tetrafe(mesh, input_template, filename_out, keywords=['NSET', 'ELSET'])
INFO:root:Converting mesh with fields:
INFO:root:{'points': array([[ 1.0637777e-01, 2.7099030e+00, 1.2464654e+00],
[ 3.1638098e+00, 2.4738045e+00, 8.2634044e-01],
[-2.8007149e-04, 2.4925890e+00, 1.6835400e+00],
...,
[ 1.7855428e+00, 3.3190203e+00, 1.1995814e+00],
[ 6.9351017e-01, 3.2858081e+00, 2.6503036e+00],
[ 2.7904041e+00, 2.9363434e+00, 2.7100861e+00]], dtype=float32), 'cells': [<meshio CellBlock, type: tetra, num cells: 122964>], 'point_data': {'medit:ref': array([1, 1, 1, ..., 1, 1, 1])}, 'cell_data': {'medit:ref': array([1, 1, 1, ..., 1, 1, 1])}, 'field_data': {}, 'point_sets': {}, 'cell_sets': {}, 'gmsh_periodic': None, 'info': None}
INFO:root:Reading Abaqus template file input_templates/tmp_example02_tens_static_steel.inp
INFO:root:Data written to file /home/gianthk/PycharmProjects/CT2FE/test_data/steel_foam/B_matrix_tetraFE.inp
Solve FE model in calculix
The following section assumes that you have the Calculix solver installed and accessible with the command ccx_2.17 or for multithread option ccx_2.17_MT
The multithread option in CalculiX is activated by defining the number of threads (default=1) used for the calculation.
This is done by setting the env variable OMP_NUM_THREADS with:
export OMP_NUM_THREADS=8
# !export OMP_NUM_THREADS=8; ccx_2.17_MT "trabecular_sample_voxelfe"
Postprocessing
Convert Calculix output to Paraview
The following sections assume that you have ccx2paraview and Paraview installed and working.
For more info visit:
https://www.paraview.org/
https://github.com/calculix/ccx2paraview
filename_out_base, ext_out = os.path.splitext(filename_out)
ccx2paraview.Converter(filename_out_base + '.frd', ['vtk']).run()
Visualize results in Paraview
!paraview filename_out_base + '.vtk'

Generate tetrahedra-FE model for non-linear analysis
input_template = "./../../input_templates/tmp_example02_tens_Nlgeom_steel.inp"
Inspect .INP template file for non-linear analysis
!cat {input_template}
** -----------------`----------------------------------
** Material definition:
*MATERIAL, NAME=STEEL_4301
*ELASTIC
210000, 0.3333333333, 0
*DENSITY
7.85e-9
*PLASTIC
2000, 0, 0
3000, 1, 0
*SOLID SECTION, ELSET=SET1, MATERIAL=STEEL_4301
** ---------------------------------------------------
** Analysis definition:
*TIME POINTS, NAME=t10, GENERATE
0, 1, 0.05
*STEP, AMPLITUDE=RAMP, UNSYMM=YES, NLGEOM=YES
*STATIC
0.2, 1, 1e-04, 0.2
** All displacements fixed at bottom:
*BOUNDARY, TYPE=DISPLACEMENT
NODES_B, 1, 3, 0
** Vertical displacement imposed at top:
*BOUNDARY, TYPE=DISPLACEMENT
NODES_T, 3, 3, +0.5
** ---------------------------------------------------
** Output request:
*EL FILE, OUTPUT=2D, TIME POINTS=t10
S,PEEQ
*NODE FILE, OUTPUT=2D
U
*NODE PRINT, TOTALS=ONLY, NSET=NODES_B
RF
*END STEP
filename_out = './../../test_data/steel_foam/B_matrix_tetraFE_Nlgeom.inp'
ciclope.mesh2tetrafe(mesh, input_template, filename_out, keywords=['NSET', 'ELSET'])
Generate CalculiX FE input file
filename_out_base, ext_out = os.path.splitext(filename_out)
ccx2paraview.Converter(filename_out_base + '.frd', ['vtk']).run()
INFO:root:Reading B_matrix_tetraFE_Nlgeom.frd
INFO:root:38341 nodes
INFO:root:122964 cells
INFO:root:Step 1, time 5.00e-02, U, 3 components, 38341 values
INFO:root:Step 1, time 5.00e-02, S, 6 components, 38341 values
INFO:root:Step 1, time 5.00e-02, PEEQ, 1 components, 38341 values
INFO:root:Step 1, time 5.00e-02, ERROR, 1 components, 38341 values
INFO:root:Step 2, time 1.00e-01, U, 3 components, 38341 values
INFO:root:Step 2, time 1.00e-01, S, 6 components, 38341 values
INFO:root:Step 2, time 1.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 2, time 1.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 3, time 1.50e-01, U, 3 components, 38341 values
INFO:root:Step 3, time 1.50e-01, S, 6 components, 38341 values
INFO:root:Step 3, time 1.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 3, time 1.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 4, time 2.00e-01, U, 3 components, 38341 values
INFO:root:Step 4, time 2.00e-01, S, 6 components, 38341 values
INFO:root:Step 4, time 2.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 4, time 2.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 5, time 2.50e-01, U, 3 components, 38341 values
INFO:root:Step 5, time 2.50e-01, S, 6 components, 38341 values
INFO:root:Step 5, time 2.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 5, time 2.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 6, time 3.00e-01, U, 3 components, 38341 values
INFO:root:Step 6, time 3.00e-01, S, 6 components, 38341 values
INFO:root:Step 6, time 3.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 6, time 3.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 7, time 3.50e-01, U, 3 components, 38341 values
INFO:root:Step 7, time 3.50e-01, S, 6 components, 38341 values
INFO:root:Step 7, time 3.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 7, time 3.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 8, time 4.00e-01, U, 3 components, 38341 values
INFO:root:Step 8, time 4.00e-01, S, 6 components, 38341 values
INFO:root:Step 8, time 4.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 8, time 4.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 9, time 4.50e-01, U, 3 components, 38341 values
INFO:root:Step 9, time 4.50e-01, S, 6 components, 38341 values
INFO:root:Step 9, time 4.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 9, time 4.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 10, time 5.00e-01, U, 3 components, 38341 values
INFO:root:Step 10, time 5.00e-01, S, 6 components, 38341 values
INFO:root:Step 10, time 5.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 10, time 5.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 11, time 5.50e-01, U, 3 components, 38341 values
INFO:root:Step 11, time 5.50e-01, S, 6 components, 38341 values
INFO:root:Step 11, time 5.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 11, time 5.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 12, time 6.00e-01, U, 3 components, 38341 values
INFO:root:Step 12, time 6.00e-01, S, 6 components, 38341 values
INFO:root:Step 12, time 6.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 12, time 6.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 13, time 6.50e-01, U, 3 components, 38341 values
INFO:root:Step 13, time 6.50e-01, S, 6 components, 38341 values
INFO:root:Step 13, time 6.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 13, time 6.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 14, time 7.00e-01, U, 3 components, 38341 values
INFO:root:Step 14, time 7.00e-01, S, 6 components, 38341 values
INFO:root:Step 14, time 7.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 14, time 7.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 15, time 7.50e-01, U, 3 components, 38341 values
INFO:root:Step 15, time 7.50e-01, S, 6 components, 38341 values
INFO:root:Step 15, time 7.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 15, time 7.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 16, time 8.00e-01, U, 3 components, 38341 values
INFO:root:Step 16, time 8.00e-01, S, 6 components, 38341 values
INFO:root:Step 16, time 8.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 16, time 8.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 17, time 8.50e-01, U, 3 components, 38341 values
INFO:root:Step 17, time 8.50e-01, S, 6 components, 38341 values
INFO:root:Step 17, time 8.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 17, time 8.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 18, time 9.00e-01, U, 3 components, 38341 values
INFO:root:Step 18, time 9.00e-01, S, 6 components, 38341 values
INFO:root:Step 18, time 9.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 18, time 9.00e-01, ERROR, 1 components, 38341 values
INFO:root:Step 19, time 9.50e-01, U, 3 components, 38341 values
INFO:root:Step 19, time 9.50e-01, S, 6 components, 38341 values
INFO:root:Step 19, time 9.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 19, time 9.50e-01, ERROR, 1 components, 38341 values
INFO:root:Step 20, time 1.0, U, 3 components, 38341 values
INFO:root:Step 20, time 1.0, S, 6 components, 38341 values
INFO:root:Step 20, time 1.0, PEEQ, 1 components, 38341 values
INFO:root:Step 20, time 1.0, ERROR, 1 components, 38341 values
INFO:root:20 time increments
INFO:root:Writing B_matrix_tetraFE_Nlgeom.01.vtk
INFO:root:Step 1, time 5.00e-02, U, 3 components, 38341 values
INFO:root:Step 1, time 5.00e-02, S, 6 components, 38341 values
INFO:root:Step 1, time 5.00e-02, S_Mises, 1 components, 38341 values
INFO:root:Step 1, time 5.00e-02, S_Principal, 3 components, 38341 values
INFO:root:Step 1, time 5.00e-02, PEEQ, 1 components, 38341 values
INFO:root:Step 1, time 5.00e-02, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.02.vtk
INFO:root:Step 2, time 1.00e-01, U, 3 components, 38341 values
INFO:root:Step 2, time 1.00e-01, S, 6 components, 38341 values
INFO:root:Step 2, time 1.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 2, time 1.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 2, time 1.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 2, time 1.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.03.vtk
INFO:root:Step 3, time 1.50e-01, U, 3 components, 38341 values
INFO:root:Step 3, time 1.50e-01, S, 6 components, 38341 values
INFO:root:Step 3, time 1.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 3, time 1.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 3, time 1.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 3, time 1.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.04.vtk
INFO:root:Step 4, time 2.00e-01, U, 3 components, 38341 values
INFO:root:Step 4, time 2.00e-01, S, 6 components, 38341 values
INFO:root:Step 4, time 2.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 4, time 2.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 4, time 2.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 4, time 2.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.05.vtk
INFO:root:Step 5, time 2.50e-01, U, 3 components, 38341 values
INFO:root:Step 5, time 2.50e-01, S, 6 components, 38341 values
INFO:root:Step 5, time 2.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 5, time 2.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 5, time 2.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 5, time 2.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.06.vtk
INFO:root:Step 6, time 3.00e-01, U, 3 components, 38341 values
INFO:root:Step 6, time 3.00e-01, S, 6 components, 38341 values
INFO:root:Step 6, time 3.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 6, time 3.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 6, time 3.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 6, time 3.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.07.vtk
INFO:root:Step 7, time 3.50e-01, U, 3 components, 38341 values
INFO:root:Step 7, time 3.50e-01, S, 6 components, 38341 values
INFO:root:Step 7, time 3.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 7, time 3.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 7, time 3.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 7, time 3.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.08.vtk
INFO:root:Step 8, time 4.00e-01, U, 3 components, 38341 values
INFO:root:Step 8, time 4.00e-01, S, 6 components, 38341 values
INFO:root:Step 8, time 4.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 8, time 4.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 8, time 4.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 8, time 4.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.09.vtk
INFO:root:Step 9, time 4.50e-01, U, 3 components, 38341 values
INFO:root:Step 9, time 4.50e-01, S, 6 components, 38341 values
INFO:root:Step 9, time 4.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 9, time 4.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 9, time 4.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 9, time 4.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.10.vtk
INFO:root:Step 10, time 5.00e-01, U, 3 components, 38341 values
INFO:root:Step 10, time 5.00e-01, S, 6 components, 38341 values
INFO:root:Step 10, time 5.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 10, time 5.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 10, time 5.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 10, time 5.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.11.vtk
INFO:root:Step 11, time 5.50e-01, U, 3 components, 38341 values
INFO:root:Step 11, time 5.50e-01, S, 6 components, 38341 values
INFO:root:Step 11, time 5.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 11, time 5.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 11, time 5.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 11, time 5.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.12.vtk
INFO:root:Step 12, time 6.00e-01, U, 3 components, 38341 values
INFO:root:Step 12, time 6.00e-01, S, 6 components, 38341 values
INFO:root:Step 12, time 6.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 12, time 6.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 12, time 6.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 12, time 6.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.13.vtk
INFO:root:Step 13, time 6.50e-01, U, 3 components, 38341 values
INFO:root:Step 13, time 6.50e-01, S, 6 components, 38341 values
INFO:root:Step 13, time 6.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 13, time 6.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 13, time 6.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 13, time 6.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.14.vtk
INFO:root:Step 14, time 7.00e-01, U, 3 components, 38341 values
INFO:root:Step 14, time 7.00e-01, S, 6 components, 38341 values
INFO:root:Step 14, time 7.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 14, time 7.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 14, time 7.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 14, time 7.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.15.vtk
INFO:root:Step 15, time 7.50e-01, U, 3 components, 38341 values
INFO:root:Step 15, time 7.50e-01, S, 6 components, 38341 values
INFO:root:Step 15, time 7.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 15, time 7.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 15, time 7.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 15, time 7.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.16.vtk
INFO:root:Step 16, time 8.00e-01, U, 3 components, 38341 values
INFO:root:Step 16, time 8.00e-01, S, 6 components, 38341 values
INFO:root:Step 16, time 8.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 16, time 8.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 16, time 8.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 16, time 8.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.17.vtk
INFO:root:Step 17, time 8.50e-01, U, 3 components, 38341 values
INFO:root:Step 17, time 8.50e-01, S, 6 components, 38341 values
INFO:root:Step 17, time 8.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 17, time 8.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 17, time 8.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 17, time 8.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.18.vtk
INFO:root:Step 18, time 9.00e-01, U, 3 components, 38341 values
INFO:root:Step 18, time 9.00e-01, S, 6 components, 38341 values
INFO:root:Step 18, time 9.00e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 18, time 9.00e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 18, time 9.00e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 18, time 9.00e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.19.vtk
INFO:root:Step 19, time 9.50e-01, U, 3 components, 38341 values
INFO:root:Step 19, time 9.50e-01, S, 6 components, 38341 values
INFO:root:Step 19, time 9.50e-01, S_Mises, 1 components, 38341 values
INFO:root:Step 19, time 9.50e-01, S_Principal, 3 components, 38341 values
INFO:root:Step 19, time 9.50e-01, PEEQ, 1 components, 38341 values
INFO:root:Step 19, time 9.50e-01, ERROR, 1 components, 38341 values
INFO:root:Writing B_matrix_tetraFE_Nlgeom.20.vtk
INFO:root:Step 20, time 1.0, U, 3 components, 38341 values
INFO:root:Step 20, time 1.0, S, 6 components, 38341 values
INFO:root:Step 20, time 1.0, S_Mises, 1 components, 38341 values
INFO:root:Step 20, time 1.0, S_Principal, 3 components, 38341 values
INFO:root:Step 20, time 1.0, PEEQ, 1 components, 38341 values
INFO:root:Step 20, time 1.0, ERROR, 1 components, 38341 values
Equivalent Plastic Strain (PEEQ) animation created in Paraview:

Post-process non-linear FE analysis results
We use the helper script dat2txt.py from Martin Kraska to clean and convert the CalculiX FE output .DAT file to a text file containing only data:
filename_dat = './../../test_data/steel_foam/B_matrix_tetraFE_Nlgeom.dat'
dat2txt.dat2txt(filename_dat, True)
INFO:root:Reading file /home/gianthk/PycharmProjects/CT2FE/test_data/steel_foam/B_matrix_tetraFE_Nlgeom.dat.
INFO:root:Written total force fx,fy,fz for group NODES_B to file: total force fx,fy,fz_NODES_B.txt.
Total time points: 59.
('total force fx,fy,fz', 'NODES_B', 59)
Load and plot results with pandas
%matplotlib inline
header = ['time', 'fx', 'fy', 'fz']
filename_txt = './../../test_data/steel_foam/total force fx,fy,fz_NODES_B.txt'
data = pd.read_table(filename_txt, delim_whitespace=True, names=header, index_col=0)
Have a look at the data produced by CalculiX
data
| fx | fy | fz | |
|---|---|---|---|
| time | |||
| 0.05 | 6.246617e-09 | -9.647296e-10 | -63.92774 |
| 0.10 | 1.269526e-10 | 2.538340e-10 | -97.62540 |
| 0.15 | 3.104711e-09 | 1.066627e-09 | -116.32770 |
| 0.20 | 9.618676e-10 | -4.711595e-10 | -129.88530 |
| 0.25 | -4.597598e-07 | -8.556020e-08 | -141.77580 |
| 0.30 | 3.020426e-07 | 3.076565e-07 | -152.58380 |
| 0.35 | -4.586224e-09 | 1.690051e-08 | -161.88950 |
| 0.40 | 1.673068e-09 | 2.573413e-09 | -168.59850 |
| 0.45 | 3.774273e-07 | 1.899347e-07 | -174.08460 |
| 0.50 | -1.608090e-07 | -1.298831e-07 | -177.40930 |
| 0.55 | 7.836687e-09 | 1.665111e-10 | -178.39990 |
| 0.60 | -9.815839e-09 | -6.166009e-09 | -179.22490 |
| 0.65 | 6.769719e-09 | 9.593283e-09 | -178.20100 |
| 0.70 | -1.156836e-09 | 1.002093e-08 | -176.15900 |
| 0.75 | -7.059488e-07 | -2.223455e-07 | -173.43490 |
| 0.80 | -1.720688e-07 | -9.126840e-08 | -170.56940 |
| 0.85 | -9.886431e-09 | -6.904714e-09 | -167.71290 |
| 0.90 | -2.093420e-08 | -4.455112e-08 | -165.11650 |
| 0.95 | -7.948244e-09 | -5.174397e-09 | -162.54210 |
| 1.00 | -8.049240e-08 | -8.431905e-08 | -160.17860 |
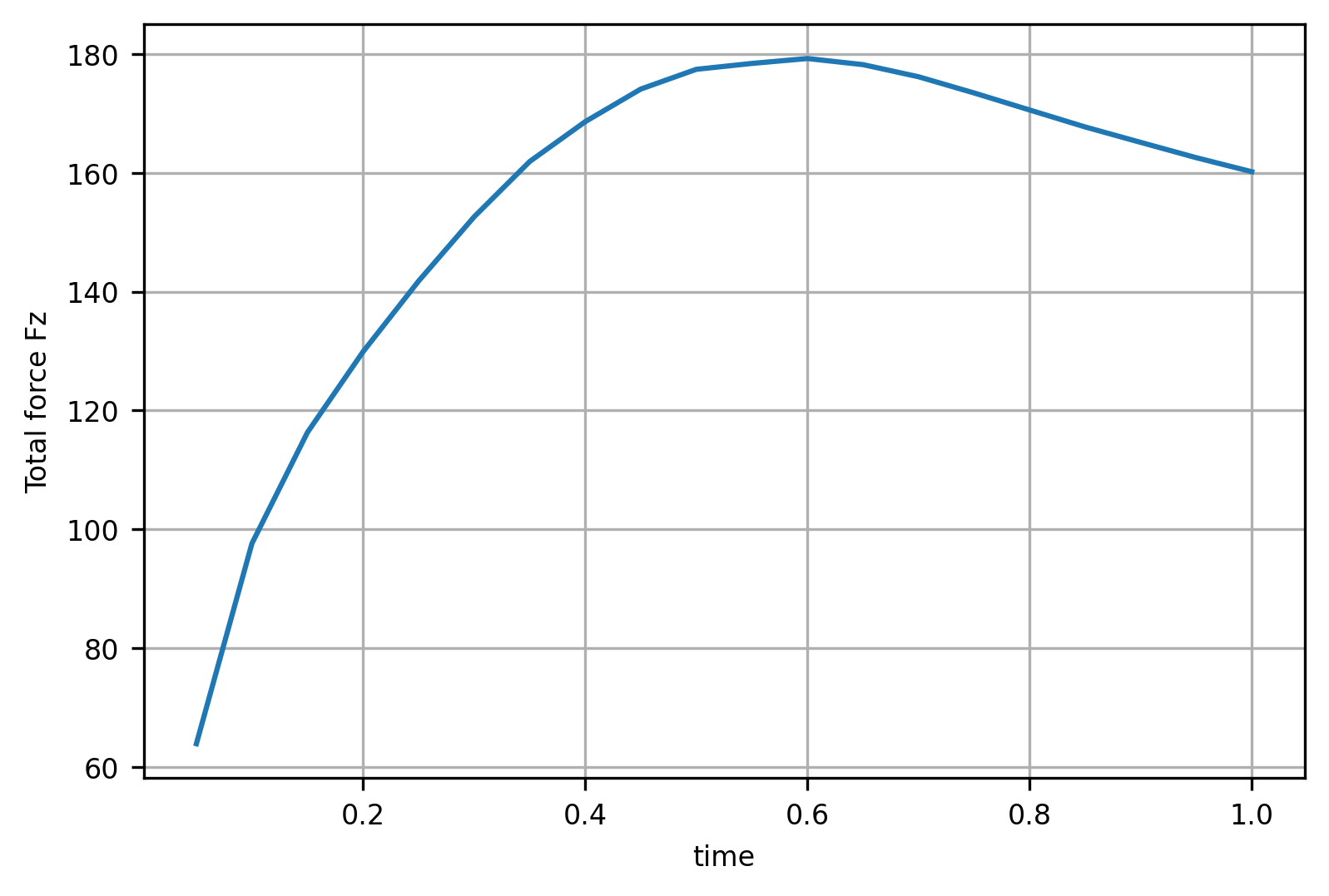
Plot Force-Displacement curve
fig, axs = plt.subplots()
data['fz'].abs().plot(ax=axs, grid=True)
axs.set_ylabel("Total force Fz")
# fig.savefig("no2_concentrations.png")
Text(0, 0.5, 'Total force Fz')